One of the biggest challenges with rare, especially ultra-rare, diseases is getting patients to the right diagnosis faster. Patients who have not yet been diagnosed or have been misdiagnosed and, as a result, are not receiving specific, targeted treatment.
About eight years ago – before AI models and large language models (LLMs) of today’s quality were available – I had the chance to explore the possibility of using artificial intelligence to identify patients suffering from a rare disease (and not being aware of it) based on health records (EHR). However, I believe that back then, we were just too early, and there wasn’t the acceptance and openness to using AI that we are facing in 2026.
So, when I took the MIT Sloan course “AI in Healthcare” earlier this year, I recalled the idea and reworked the use case, drawing on the state of the art available in 2026. I am happy to share the outcome as a sketch.
Kindly consider that there is certainly more “flesh to the bone” that I generously skipped to avoid overloading this introductory overview article. It is a sketch. There are many more details to consider. And certainly, in addition, all basic and AI-agnostic best practices for successfully running healthcare projects apply.
But it is not actually rocket science, and I have personally been excited about the new opportunities while working on the case. If you would like to know more, feel free to approach me anytime, please.
The challenge
In rare diseases, many patients fall through the cracks of healthcare systems and standard diagnostic pathways. They are misdiagnosed or undiagnosed for years, cycling from specialist to specialist, exposed to unnecessary procedures, and suffering while their disease deteriorates. Today, doctors are supported by online tools that provide a “suspicion rate”, where anything above a certain threshold qualifies for a confirmatory biomarker and/or genetic test. So, if an AI could achieve higher precision and identify additional cases based on new patterns (e.g., sociodemographic features), that would be a big leap forward.
The opportunity goes beyond patients. HCPs can provide their patients with clarity more quickly and begin treatment earlier. Payers benefit from reduced diagnostic odyssey costs. Pharma industry gains access to previously invisible patient populations for their therapies, clinical trials, and real-world evidence. Patient advocacy groups get an opportunity to increase disease awareness. And patients’ families, who also carry an enormous emotional and care burden, benefit from reduced uncertainty and purposeful support.
I also see potential to move ambition, efforts, and investments from an inefficient approach today, which is creating awareness and educating thousands of clinical specialists to recognize and diagnose one single rare disease (of ~7`000), to … simply diagnosing the disease.
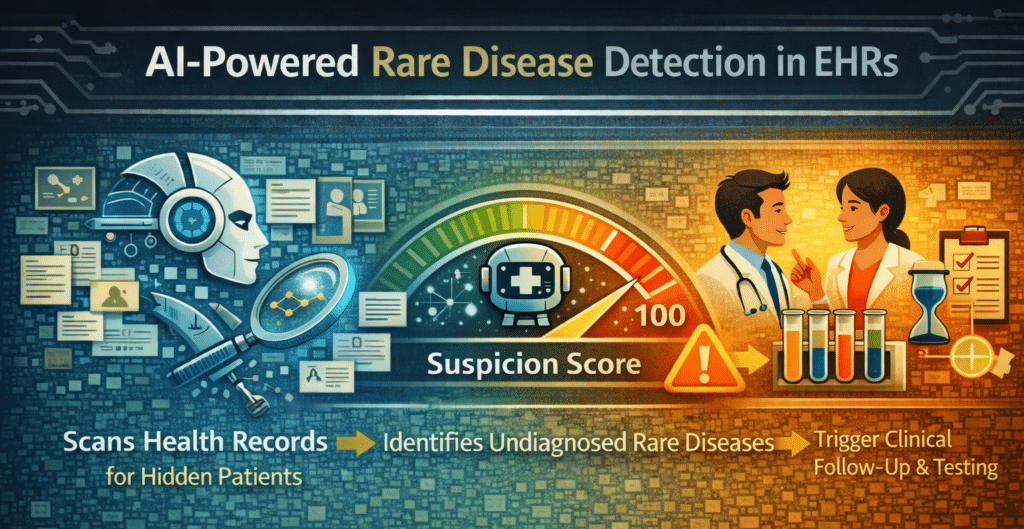

The approach
The approach is to let an AI model scan health records for signs of a specific rare disease, which might be specific symptom combinations and observations across medical specialties as well as new patterns self-learned by the model.
The AI output is a likelihood or ‘suspicion’ score, not a diagnosis. Above a defined confidence threshold, it triggers confirmatory biomarker and genetic testing. So, AI output is the beginning of a clinical decision & care chain, rather than an endpoint.
The task is a typical pattern-recognition problem across large, heterogeneous, poorly structured datasets, with data scattered across medical specialties. This is where AI (specifically NLP-based approaches) can outperform manual clinical review. AI can process records at the population level, detect feature combinations that very few clinicians can, and do so unbiasedly and continuously.
But the AI’s suitability and all the potential benefits mentioned before are conditional. They depend on (at least) …
- an agreed governance model & clear accountabilities across institutions and contributors involved,
- regulatory compliance,
- accessible, high-quality data,
- the model’s accuracy,
- actionable metrics,
- smooth clinical workflow integration,
- interpretability/explainability of the model’s output,
- and, as a result, on the trust and confidence of all stakeholders.
Data might be one of the biggest limitations. Rare diseases mean low volumes; multinational aggregation is likely helpful; data quality is ‘mixed’ with inconsistencies across and within countries, data structures changing over time, and completeness is more an aspiration than a reality. Data access is country- and jurisdiction-dependent; requirements under divergent regulatory frameworks must be met proactively (FDA, EMA, CSV, MDR, HIPAA, national data privacy laws). As an example, federated learning can address cross-border data transfer restrictions, but at the cost of increased regulatory complexity. Same with upscaling, which will additionally introduce distributional shifts, necessitating robust assessment followed by continuous monitoring. Therefore, the framework advocates deliberate exclusion of countries where conditions are not (sufficiently) met.
Data quality monitoring must be automated via a statistical process control system that flags quality issues and distributional shifts and halts predictions rather than letting the model adapt autonomously. Patient registries can serve as validation data, but might be too limited in scope for primary training.
On accuracy: I would not like to put higher expectations on AI than on standard clinical practice. Achieving anything above the standard clinical success rate, plus broader coverage, is already significant. The failure mode to avoid is false negatives (missed patients who continue to suffer). False positives create patient stress (anxiety, invasive testing), but at the end, a negative result also helps exclude a disease in the diagnostic process, which is often appreciated. Copy-forwarding risks and the needle-in-a-haystack problem are explicitly named: a single buried symptom mention might be the key clue, and the AI must reliably find it.
The AI must cite which clinical features from which records led to each flag, along with the score. Interpretability is non-negotiable. When the AI flags a patient, it must show which clinical features from which records led to that flag. This is a direct application of the course’s emphasis on transparency and citation-backed outputs. Limiting interpretability to technical AI metrics would miss the point for key stakeholders. The distinction between extractive AI (transparent, traceable) and generative AI (risk of hallucinations) must be maintained throughout the architecture. A particular challenge could be that, if the AI identifies patients using non-clinical features (sociodemographic data, lifestyle patterns), clinicians might resist what they cannot explain. By the way, interpretability requirements differ by stakeholder: HCPs demand scientific evidence; patients need lay-language output and empathy; payers and regulators require compliance and proof of the system’s effectiveness, reliability, and safety.
Clinical workflow integration is where many promising digital health approaches have failed in the past, not because the technology was bad, but because the follow-up process was not thought through. Governance, seamless integration of all contributors and stakeholders, as well as clear & agreed accountabilities, are critical to the approach. The clinical implementation plan must be developed alongside the AI. How can an assigned physician be identified? How can the physician be motivated to run a confirmatory lab test? How does this fit into the national reimbursement scheme? These are not afterthoughts. And honestly, the smartest AI outcome is useless if the subsequent clinical workflow is not smart, effective, and robust as well. Or as I would paraphrase it: “Human and AI hand in hand”.
The architecture
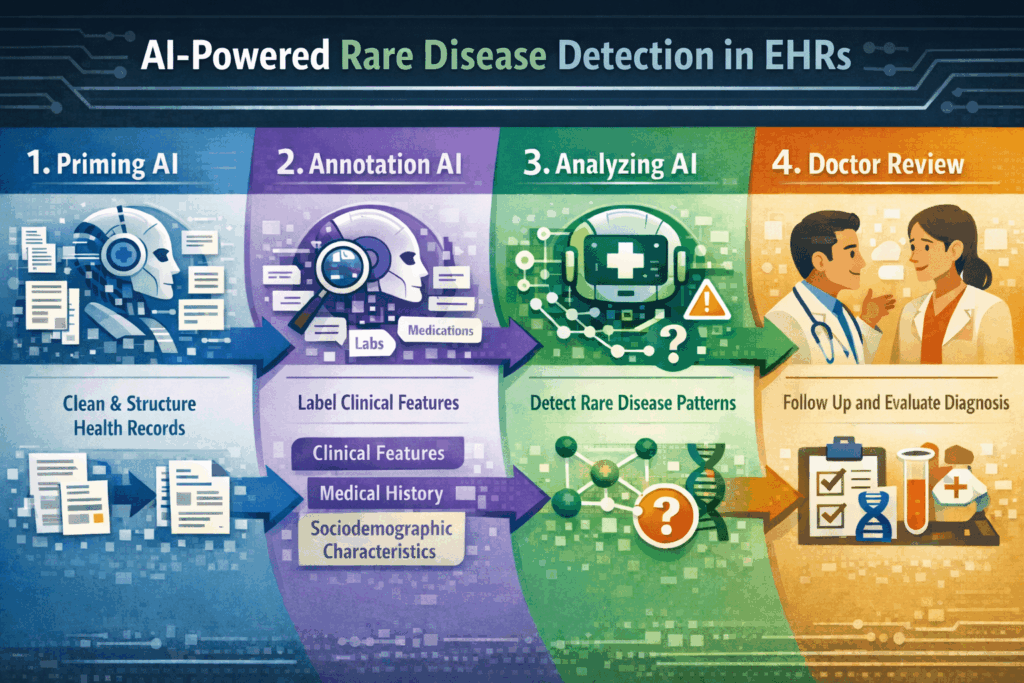
I imagine a multi-stage AI architecture:
- First, a priming AI (extractive, NLP-based) that consolidates, cleans, curates, and structures messy and chaotic health records. This AI focuses on document and data structure, not clinical content, so it can be trained on a much wider, disease-agnostic dataset.
- Second, an annotation-support AI that assists with labeling clinical features, potentially leveraging zero-shot or few-shot extraction from large language models to mitigate data scarcity.
- Third, an AI that analyzes data to identify candidate patients by matching clinical feature combinations to known diagnostic patterns (“tree-based”); it is lean, simple, close to general diagnostic concepts physicians are used to, and interpretability seems built in. And, over time, possibly discovering novel correlations beyond established clinical knowledge.
- Fourth, human doctors follow up and make decisions based on the AI’s findings. Like, informing the patient. Or deciding the next best confirmatory diagnostic step? This will require sufficient ‘interpretability‘ of AI output (allowing the HCP to understand how the AI came to its conclusion). But also a smooth integration into clinical workflows and a smart interface between AI and humans working hand in hand (I am a strong advocate of the ‘human+AI collective intelligence‘ concept).
A multi-stage architecture aligns with a preference for a lean initial setup and supports federated learning at the same time.
A couple of best practises can be implemented by design. Including citations back to source material is directly applicable. Zero-shot and few-shot extraction capabilities of LLMs can extract symptoms and clinical features without disease-specific training data, and are a potential game-changer for rare diseases. The distinction between extractive and generative AI is deliberate: extractive is more transparent, traceable, and free of hallucinations. Making this distinction additionally supports interpretability and regulatory defensibility.
Proposal for effective KPIs for the AI approach
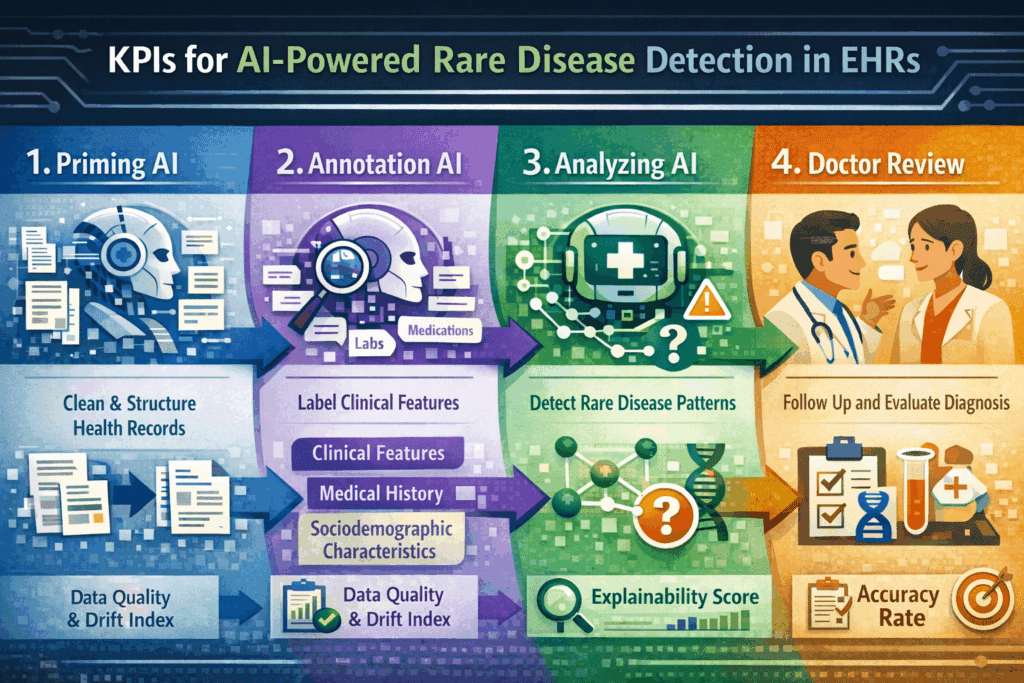
- Accuracy Rate. Measures the proportion of AI-flagged patients subsequently confirmed (positive predictive value) and the proportion of true cases identified (sensitivity). Target: sensitivity and an acceptable false-positive rate within a threshold to be agreed with clinical stakeholders. Monitored continuously against confirmatory test outcomes. Degradation triggers model revalidation.
- Data Quality and Drift Index. Tracks the stability of input data distribution, especially every time an additional source (country, EHR system) gets connected. An automated process monitors feature distributions, data completeness, and structural consistencies. This operationalizes the concern about silent failures from data quality and drift issues, and provides auditable evidence of data governance for regulatory compliance.
- Explainability Score. Percentage of AI flags accompanied by a complete, traceable citation chain linking the flag to specific features in specific source records. Target: 100%. Periodic stakeholder audits assess whether HCPs rate explanations as scientifically sufficient and whether patient-facing outputs provide high customer experience scores. This adds to the documentation for MDR and CSV/CSA compliance, where traceability of AI decision-making is a formal requirement.
Guiding questions for an effective setup
- What regulatory pathway applies in each target country, considering the full process from data access to clinical integration?
- What volume and quality of accessible, well-annotated patient-record data exist per target country?
- How will the extractive priming AI handle data structure heterogeneity across institutions, countries, and time periods without introducing systematic errors?
- What is the clinically acceptable confidence threshold, relative to false-positive burden and false-negative risk?
- How will the AI provide traceable, understandable, citation-backed explanations for each flag?
- Which pre-trained foundational models are being evaluated, and what domain-specific validation confirms suitability for rare disease NLP with limited labeled data?
- How will continuous monitoring detect distributional shifts and model drift, and what are the halt criteria?
- How will AI-discovered novel diagnostic patterns beyond established clinical knowledge be validated and made clinically acceptable?
- How does the end-to-end clinical workflow operate: who receives the flag, who decides, who acts, who informs the patient, and what safeguards prevent flags from being ignored?
- Which human-machine interfaces are required, and how can they provide excellent UX?
Final words
In 2026, AI technology is much more advanced than it was in 2018. In my opinion, the AI approach to identifying hidden cases of rare diseases in health records can work. But the AI’s suitability and all the potential benefits mentioned before are conditional. They depend on accessible, high-quality data, on the model’s accuracy, on clinical workflow integration, on regulatory compliance, on the interpretability/explainability of the model’s output, and, as a result, on the trust and confidence of all stakeholders.
But mostly with messy raw data, with a compliance-ready design, and last but not least with clinical acceptance and smooth integration into clinical workflows. But I think the approach sketched has shown that by actively considering and tackling those challenges, there should be no more excuses for not providing a clear diagnosis to rare disease patients and their families earlier.



